Tropical Plant Research
An International Journal by Society for Tropical Plant Research
2018, VOLUME 5 ISSUE 2Pages: 250-259
Assessment of genetic diversity in natural populations of Calamus guruba Buch.-Ham. ex Mart. using ISSR marker
Rajendra Kumar Meena*, Hans Raj, Pratibha Sharma, Sandeep Yadav, Rama Kant and Maneesh Singh Bhandari
*Division of Genetics & Tree Propagation, Forest Research Institute, Dehradun-248195, Uttarakhand, India
Abstract:
Rattan palms are one of the important nontimber forest produce which forms an integral part of the tribal population and contribute significantly in the rural economy. The demand of the rattan resources is substantial but most of the raw material is being collected from natural stands; and therefore, there is tremendous pressure on natural populations. Calamus guruba is one of the important rattan species of India and extensively harvested for a wide range of end products. In the present study, three populations of C. guruba naturally growing in Mizoram were characterized for their genetic diversity and population structure usingInter-Simple Sequence Repeat (ISSR) markers. A total of 197 bands generated in PCR amplification of 67 individuals with 10 ISSR primers. Among all the primers UBC 808 displayed maximum 25, while UBC 807 and UBC 827 showed the lowest 13 numbers of bands. Relatively high genetic diversity (PPB = 98.98 %, h = 0.271, I = 0.341) was recorded for the species with a moderate level of genetic differentiation (PhiPT = 0.282). Analysis of molecular variance revealed that the genetic differentiation among the populations is the significant and large proportion of the genetic variation (72%) resides among the individuals within a population, whereas only 28 % were found among the populations. UPGMA tree, PCoA plot and STRUCTURE analysis revealed that two ancestral groups captured the entire genetic variability. The baseline study on C. guruba is of paramount importance in devising programs for conservation and improvement.
Rattan palms are one of the important nontimber forest produce which forms an integral part of the tribal population and contribute significantly in the rural economy. The demand of the rattan resources is substantial but most of the raw material is being collected from natural stands; and therefore, there is tremendous pressure on natural populations. Calamus guruba is one of the important rattan species of India and extensively harvested for a wide range of end products. In the present study, three populations of C. guruba naturally growing in Mizoram were characterized for their genetic diversity and population structure usingInter-Simple Sequence Repeat (ISSR) markers. A total of 197 bands generated in PCR amplification of 67 individuals with 10 ISSR primers. Among all the primers UBC 808 displayed maximum 25, while UBC 807 and UBC 827 showed the lowest 13 numbers of bands. Relatively high genetic diversity (PPB = 98.98 %, h = 0.271, I = 0.341) was recorded for the species with a moderate level of genetic differentiation (PhiPT = 0.282). Analysis of molecular variance revealed that the genetic differentiation among the populations is the significant and large proportion of the genetic variation (72%) resides among the individuals within a population, whereas only 28 % were found among the populations. UPGMA tree, PCoA plot and STRUCTURE analysis revealed that two ancestral groups captured the entire genetic variability. The baseline study on C. guruba is of paramount importance in devising programs for conservation and improvement.
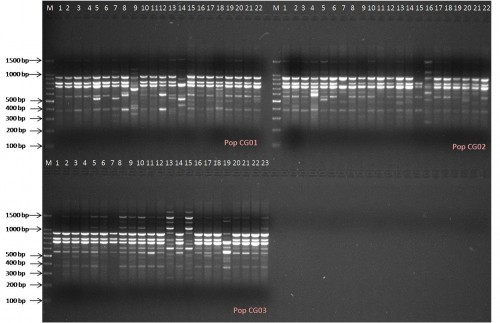
Fig.: Band profile generated by ISSR primer PCP1 in 3 populations of Calamus guruba Buch.-Ham. ex Mart.
| 0 | 1 | 2 | 5 | 9 | 3 | 7 | 2 |


