Tropical Plant Research
An International Journal by Society for Tropical Plant Research
2016, VOLUME 3 ISSUE 1Pages: 191-198
Microsatellite markers based heterozygosity assessment in Jatropha curcas L.: A potential bioenergy crop
Ramanuj Maurya* and Hemant Kumar Yadav
Abstract:
The tree breeding is more difficult by the changes that occur during the transition from juvenility to maturity. A correlation between individual heterozygosities of parents and their offspring arises from the fact that, at most allelic frequencies, heterozygous parents produce higher proportion of heterozygous progeny than do homozygous parents. Microsatellite markers are an efficient tool for the assessment of heterozygosity and homozygosity. In order to assess the level of heterozygosity of Jatropha curcas, 56 SSRs markers were used for genotyping 48 progeny derived from selfed seeds of a single J. curcas plant. Out of 56, 7 SSRs could not produce sufficient and significant data as they failed to amplify in more than 70% genotypes and thus not considered for further analysis. Therefore, genotypic data of 49 SSRs were used for heterozygosity assessment. Out of 49 SSRs, 31 SSRs were found to be monomorphic and 18 polymorphic indicating homozygosity and heterozygosity on plants, respectively. The polymorphic SSRs showed allele variation from 2 to 9 with an average of 3.56 alleles per SSRs. The SSR JGM_CD232 showed maximum of 9 alleles followed by SSR JGM_CD348, JGM_CD421, and JGM_CD092 with 5 alleles. The heterozygosity, calculated as proportion of heterozygous individuals in population, varied from 0.00 to 1.00 with an average of 0.37. However, majority of the markers (61%, 11 out of 18) showed heterozygosity variation from 0.00 to 0.22 indicating low level of heterozygosity at these loci. The rest 7 SSRs showed heterozygosity from 0.6 to 1.0 (mean 0.84) indicating higher proportion of heterozygosity at these loci. In the present investigation, the heterozygosity assessment in J. curcas indicating low level of heterozygosity showed there is need to create genetic variability in J. curcas for genetic improvement.
The tree breeding is more difficult by the changes that occur during the transition from juvenility to maturity. A correlation between individual heterozygosities of parents and their offspring arises from the fact that, at most allelic frequencies, heterozygous parents produce higher proportion of heterozygous progeny than do homozygous parents. Microsatellite markers are an efficient tool for the assessment of heterozygosity and homozygosity. In order to assess the level of heterozygosity of Jatropha curcas, 56 SSRs markers were used for genotyping 48 progeny derived from selfed seeds of a single J. curcas plant. Out of 56, 7 SSRs could not produce sufficient and significant data as they failed to amplify in more than 70% genotypes and thus not considered for further analysis. Therefore, genotypic data of 49 SSRs were used for heterozygosity assessment. Out of 49 SSRs, 31 SSRs were found to be monomorphic and 18 polymorphic indicating homozygosity and heterozygosity on plants, respectively. The polymorphic SSRs showed allele variation from 2 to 9 with an average of 3.56 alleles per SSRs. The SSR JGM_CD232 showed maximum of 9 alleles followed by SSR JGM_CD348, JGM_CD421, and JGM_CD092 with 5 alleles. The heterozygosity, calculated as proportion of heterozygous individuals in population, varied from 0.00 to 1.00 with an average of 0.37. However, majority of the markers (61%, 11 out of 18) showed heterozygosity variation from 0.00 to 0.22 indicating low level of heterozygosity at these loci. The rest 7 SSRs showed heterozygosity from 0.6 to 1.0 (mean 0.84) indicating higher proportion of heterozygosity at these loci. In the present investigation, the heterozygosity assessment in J. curcas indicating low level of heterozygosity showed there is need to create genetic variability in J. curcas for genetic improvement.
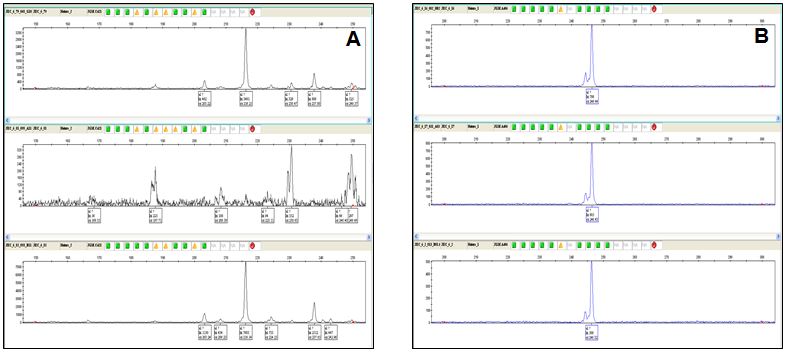
Fig.: Snap shot showing A, Polymorphic (heterozygous) B, monomorphic (homozygous) allele.
| 0 | 1 | 2 | 6 | 0 | 4 | 2 | 1 |


